NPs Basic Information
|
Name |
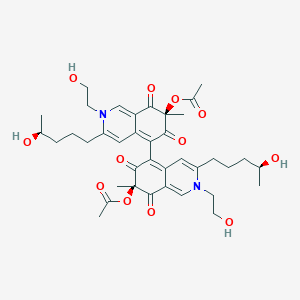
Chaetoglobin B
|
| Molecular Formula | C38H48N2O12 | |
| IUPAC Name* |
[(7S)-5-[(7S)-7-acetyloxy-2-(2-hydroxyethyl)-3-[(4S)-4-hydroxypentyl]-7-methyl-6,8-dioxoisoquinolin-5-yl]-2-(2-hydroxyethyl)-3-[(4S)-4-hydroxypentyl]-7-methyl-6,8-dioxoisoquinolin-7-yl] acetate
|
|
| SMILES |
C[C@@H](CCCC1=CC2=C(C(=O)[C@](C(=O)C2=CN1CCO)(C)OC(=O)C)C3=C4C=C(N(C=C4C(=O)[C@@](C3=O)(C)OC(=O)C)CCO)CCC[C@H](C)O)O
|
|
| InChI |
InChI=1S/C38H48N2O12/c1-21(43)9-7-11-25-17-27-29(19-39(25)13-15-41)33(47)37(5,51-23(3)45)35(49)31(27)32-28-18-26(12-8-10-22(2)44)40(14-16-42)20-30(28)34(48)38(6,36(32)50)52-24(4)46/h17-22,41-44H,7-16H2,1-6H3/t21-,22-,37+,38+/m0/s1
|
|
| InChIKey |
HPLABOYPZBUPPP-QJJXLEMDSA-N
|
|
| Synonyms |
Chaetoglobin B
|
|
| CAS | NA | |
| PubChem CID | 102029057 | |
| ChEMBL ID | NA |
*Note: the IUPAC Name was collected from PubChem.
Chemical Classification: |
|
|
|---|
——————————————————————————————————————————
NPs Species Source
| Endophyte ID | Endophyte Name | Family | Genus | Taxonomy ID | GenBank ID | Closest GenBank ID | Reference | |
|---|---|---|---|---|---|---|---|---|
| Endophyte ID | Endophyte Name | Family | Genus | Taxonomy ID | GenBank ID | Closest GenBank ID | Reference |
NPs Biological Activity
| Bioactivity Name | Target ID | Target Name | Target Type | Target Organism | Target Organism ID | Potency of Bioactivity | Activity Type | Value | Unit | Endophyte ID | Endophyte Name | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Bioactivity Name | Target ID | Target Name | Target Type | Target Organism | Target Organism ID | Potency of Bioactivity | Activity Type | Value | Unit | Endophyte ID | Endophyte Name |
NPs Physi-Chem Properties
| Molecular Weight: | 724.8 | ALogp: | 0.8 |
| HBD: | 4 | HBA: | 14 |
| Rotatable Bonds: | 17 | Lipinski's rule of five: | Rejected |
| Polar Surface Area: | 208.0 | Aromatic Rings: | 4 |
| Heavy Atoms: | 52 | QED Weighted: | 0.141 |
——————————————————————————————————————————
NPs ADMET Properties*
ADMET: Absorption
| Caco-2 Permeability: | -5.483 | MDCK Permeability: | 0.00007690 |
| Pgp-inhibitor: | 0.996 | Pgp-substrate: | 0.796 |
| Human Intestinal Absorption (HIA): | 0.934 | 20% Bioavailability (F20%): | 1 |
| 30% Bioavailability (F30%): | 0.999 |
——————————————————————————————————————————
ADMET: Distribution
| Blood-Brain-Barrier Penetration (BBB): | 0.165 | Plasma Protein Binding (PPB): | 70.70% |
| Volume Distribution (VD): | 1.555 | Fu: | 13.99% |
——————————————————————————————————————————
ADMET: Metabolism
| CYP1A2-inhibitor: | 0.395 | CYP1A2-substrate: | 0.065 |
| CYP2C19-inhibitor: | 0.07 | CYP2C19-substrate: | 0.793 |
| CYP2C9-inhibitor: | 0.247 | CYP2C9-substrate: | 0.034 |
| CYP2D6-inhibitor: | 0.004 | CYP2D6-substrate: | 0.009 |
| CYP3A4-inhibitor: | 0.854 | CYP3A4-substrate: | 0.444 |
——————————————————————————————————————————
ADMET: Excretion
| Clearance (CL): | 1.561 | Half-life (T1/2): | 0.809 |
——————————————————————————————————————————
ADMET: Toxicity
| hERG Blockers: | 0.04 | Human Hepatotoxicity (H-HT): | 0.958 |
| Drug-inuced Liver Injury (DILI): | 0.954 | AMES Toxicity: | 0.399 |
| Rat Oral Acute Toxicity: | 0.004 | Maximum Recommended Daily Dose: | 0.958 |
| Skin Sensitization: | 0.865 | Carcinogencity: | 0.181 |
| Eye Corrosion: | 0.003 | Eye Irritation: | 0.005 |
| Respiratory Toxicity: | 0.846 |
——————————————————————————————————————————
*Note: the ADMET properties was calculated by ADMETlab 2.0. Reference: PMID: 33893803.
Similar Compounds*
Compounds similar to EMNPD with top10 similarity:
| Similar NPs | Similar Drugs | ||||||
|---|---|---|---|---|---|---|---|
| NPs ID | NPs 2D Structure | Similarity Score | TTD ID | Drug 2D Structure | Similarity Score | ||
| ENC004069 | 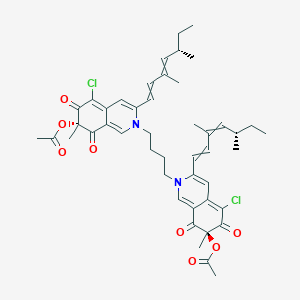 |
0.451 | D08FPM | 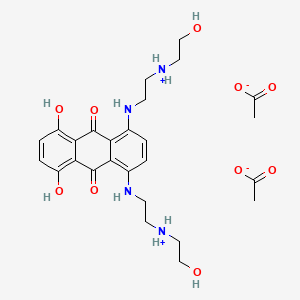 |
0.218 | ||
| ENC001870 | 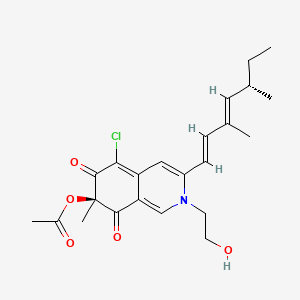 |
0.325 | D0WY9N | 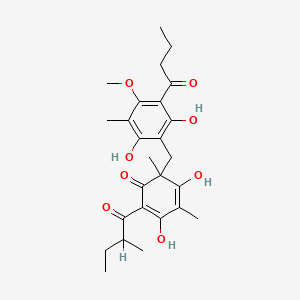 |
0.206 | ||
| ENC002463 | 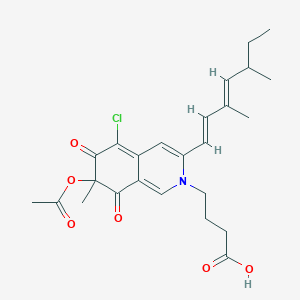 |
0.310 | D04VEJ | 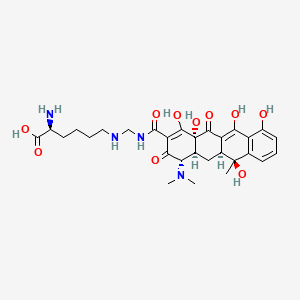 |
0.197 | ||
| ENC003605 | 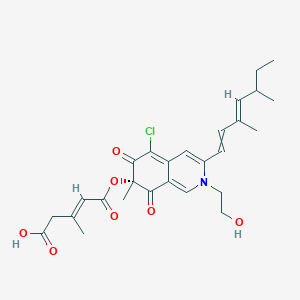 |
0.304 | D00FSV | 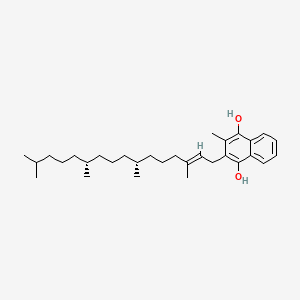 |
0.197 | ||
| ENC006052 | 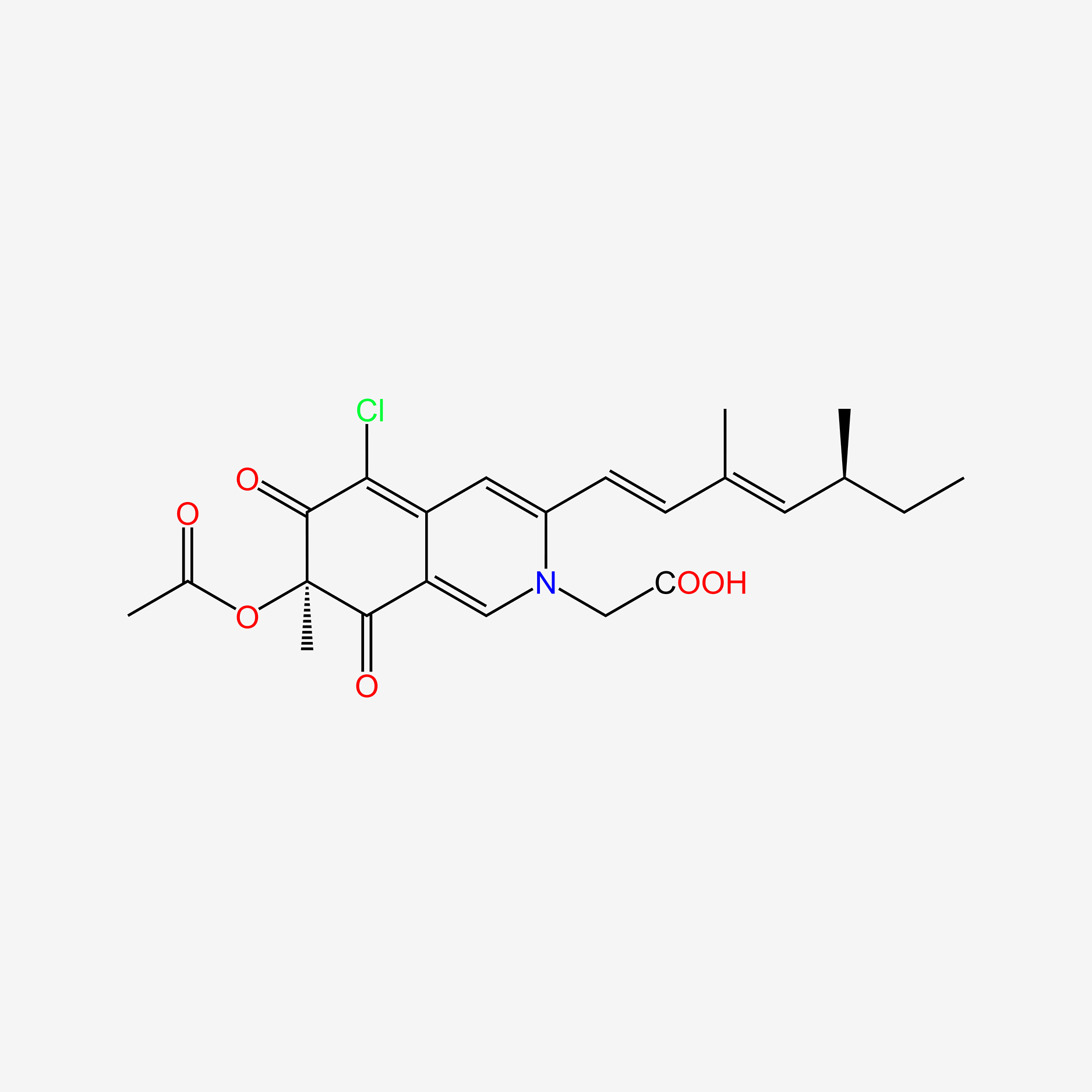 |
0.291 | D0T5XN | 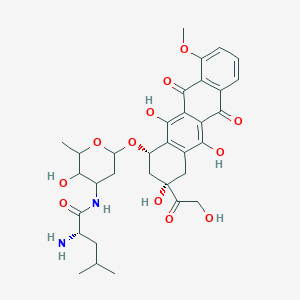 |
0.187 | ||
| ENC005593 | 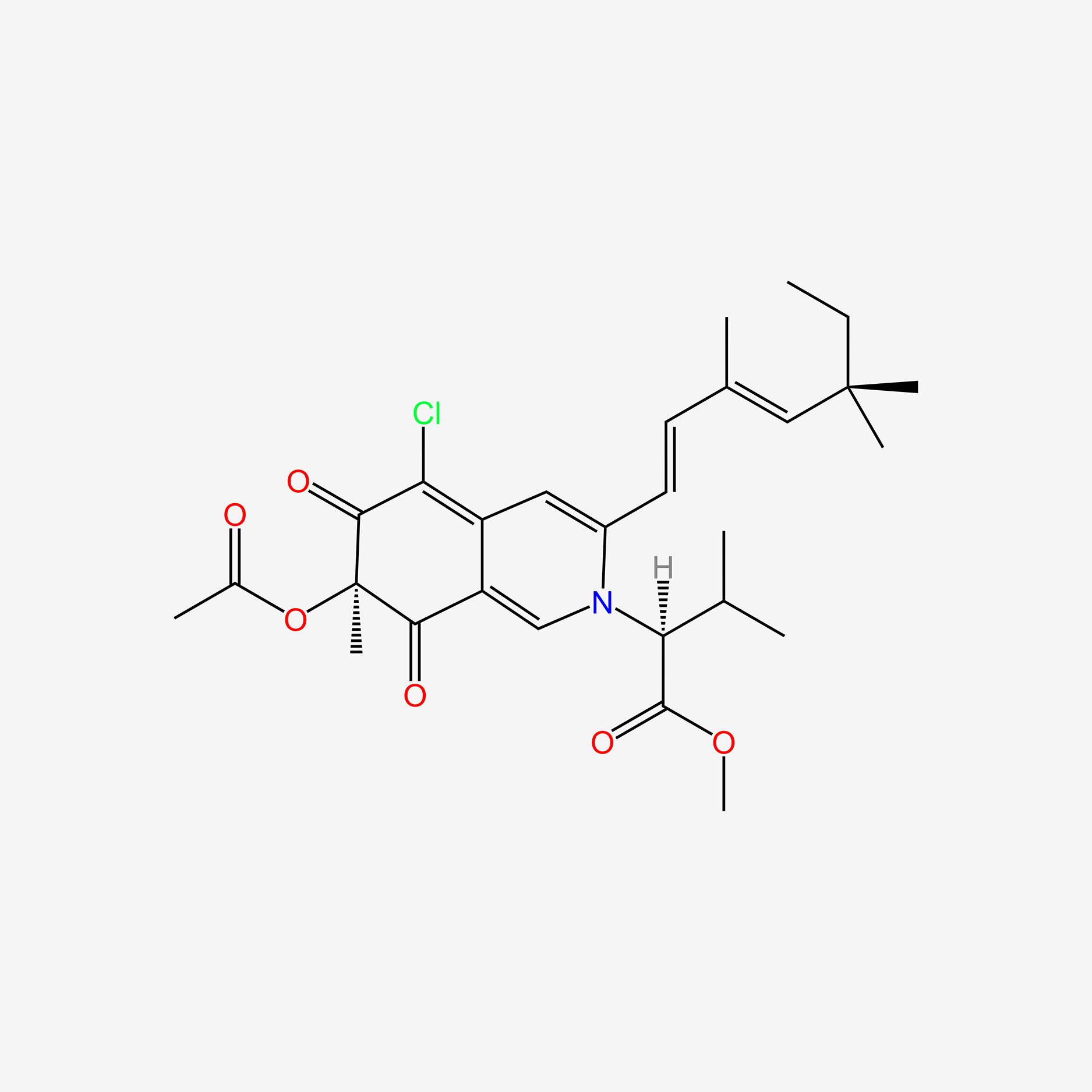 |
0.272 | D0N1FS | 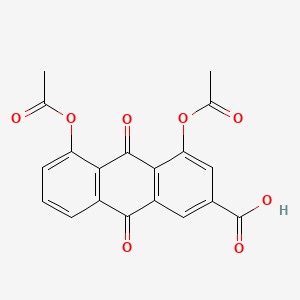 |
0.183 | ||
| ENC003676 |  |
0.265 | D00ZCN | 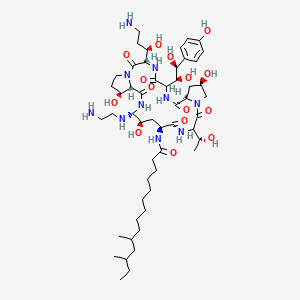 |
0.182 | ||
| ENC004761 | 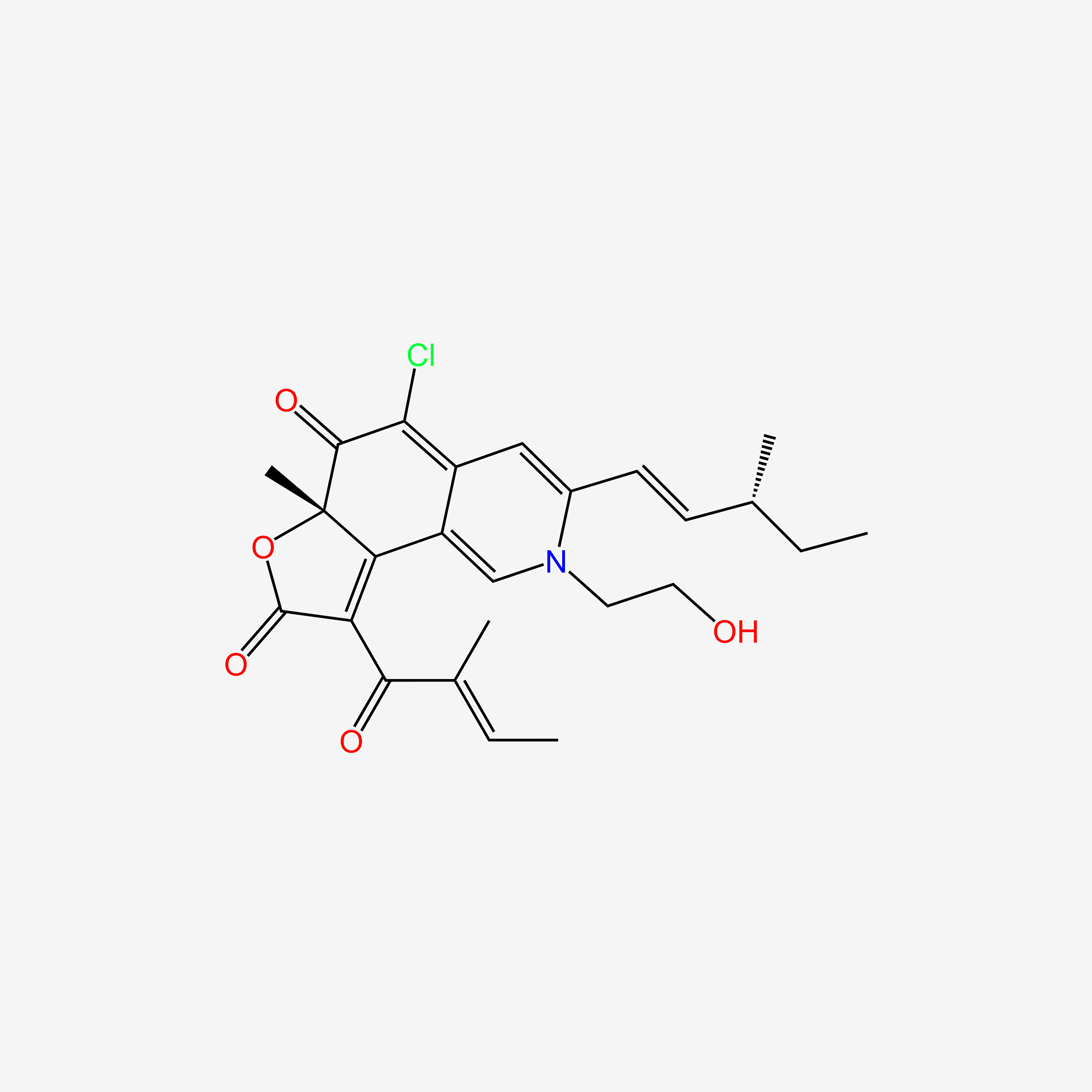 |
0.249 | D00WMK | 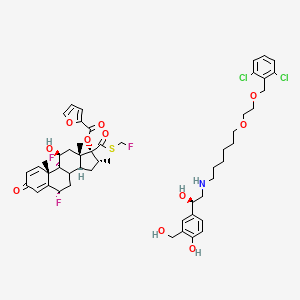 |
0.181 | ||
| ENC006054 | 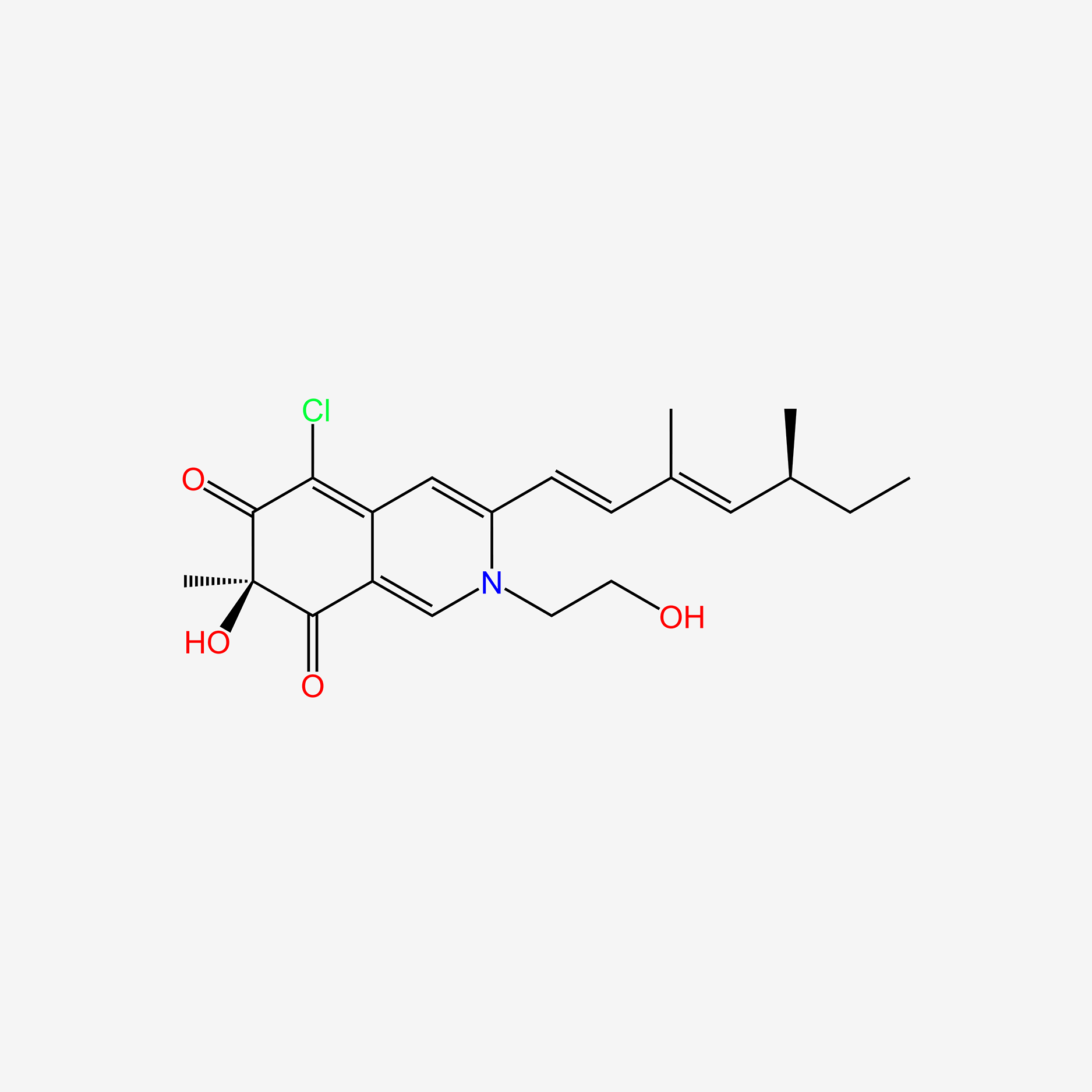 |
0.247 | D05BJD | 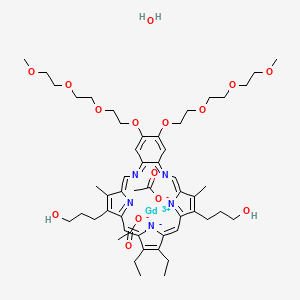 |
0.181 | ||
| ENC006053 | 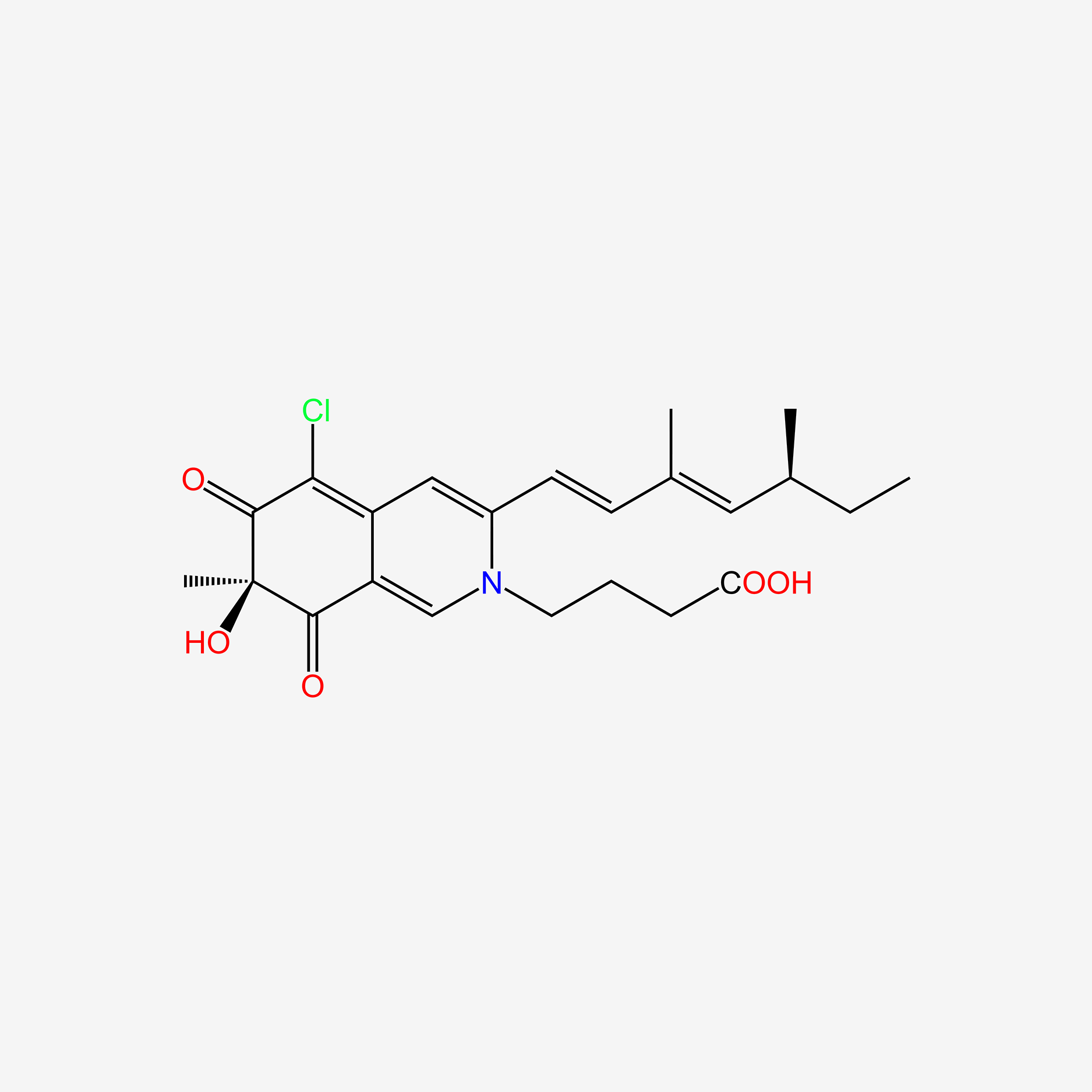 |
0.243 | D0Y4YG | 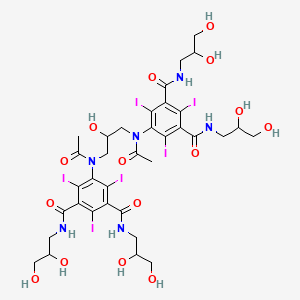 |
0.179 | ||